INTRODUCTION
Pancreatic ductal adenocarcinoma (PDAC) is the third most lethal cancer in the USA, with a 10% five-year survival rate [1]. The majority of PDAC cases are diagnosed at an advanced stage, mostly due to lack of early specific symptoms, and absence of effective screening strategies. PDAC in its early or precursor stages is potentially resectable and curable [2]. Although screening is recommended for individuals with an inherited predisposition, who comprise less than 10% of all PDAC cases [3], there is currently no screening program for the general population.
Our prior work demonstrated that leveraging Machine Learning on diagnoses from Electronic Health Records (EHRs), can identify individuals at high-risk for PDAC, up to 1 year before current diagnosis [4].
We aim to improve the performance of our model by using a multi-center dataset, and adding lab test features.
MATERIALS & METHODS
EHR data from TriNetX, a federated health research network, was utilized to develop Logistic Regression (LR) models, using diagnoses and lab test data from 32 Health Care Organizations in the US (2015-2020).
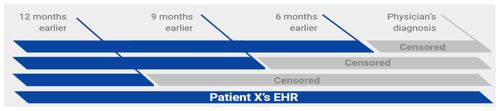
PDAC patients were identified using ICD codes, and validated with tumor registry and pathology data. Patients 60-80 years old, with one or more clinical encounter/s, minimum 6 months prior to PDAC diagnosis, were included, using prediction time cutoffs of 180, 270, 360 days before PDAC diagnosis (Fig. 1).
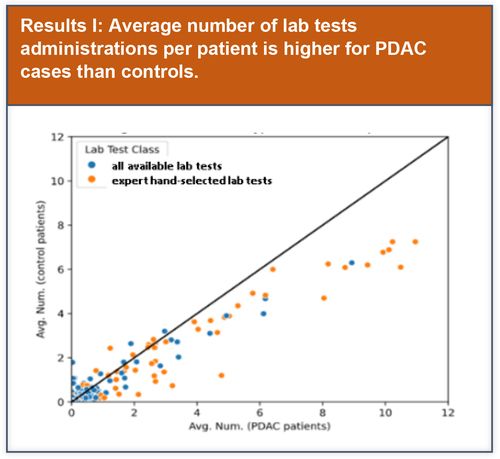
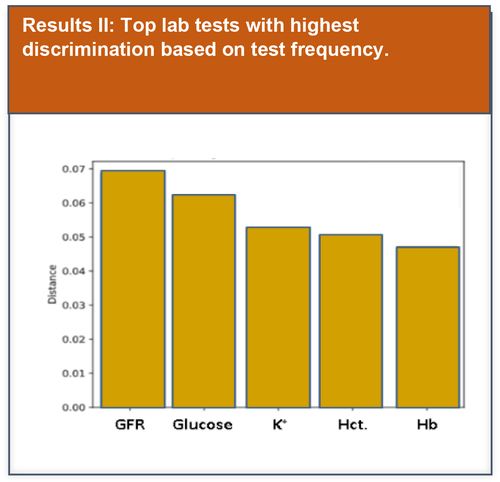
Preliminary basic data analysis was initially performed to explore potential lab test features that could be used to improve model performance.
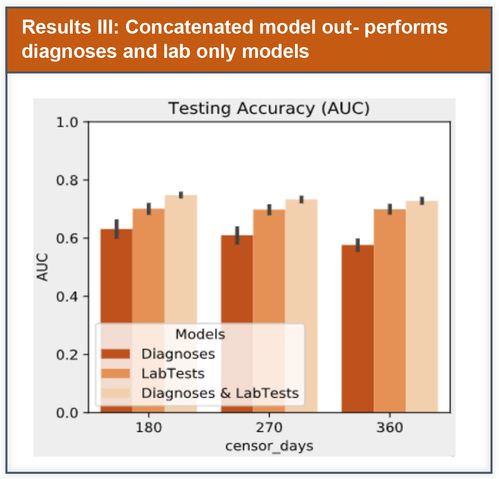
LR models were compared using Area Under the Receiver Operating Characteristic Curve (AUC), 95% Confidence Interval using empirical bootstrap over test data were computed.
We used L2-regularized LR, and performed evaluation using cross-validation. In contrast to prior published work that used predefined feature sets, we incorporated a wide range of indicators, and relied on regularization to address potential overfitting risk.